There’s two exciting news today in embryo selection. The technology is finally moving forward at an appreciable rate. Maybe we are not dysgenics doomed after all.
First off, Herasight — the new startup for embryo selection I covered in July — just published their follow-up validation study concerning the prediction of intelligence: Within-family validation of a new polygenic predictor of general cognitive ability (there’s a blogpost summary as well). Spencer Moore has a thread giving a summary of the findings. The abstract reads:
We present a polygenic score (PGS) for general cognitive ability (GCA) that demonstrates a substantial increase in predictive accuracy both among unrelated individuals and within-families relative to existing predictors. In the UK Biobank (UKB), our PGS achieves a standardized regression coefficient with fluid intelligence of β = 0.406 (SE 0.009), corresponding to an R 2 of 16.4% (95% confidence interval: [15.1%, 17.9%]). In a sample of 4,642 sibling pairs from UKB, we estimate the standardized within-family effect of the PGS to be δ = 0.355 (SE 0.0218), indicating only slight attenuation of prediction ability within-family (δ/β = 0.876, SE 0.050). We obtain similar results in a sample of 736 9 – 10-year-old European ancestry sibling pairs from the Adolescent Brain Cognitive Development (ABCD) cohort. After correcting for the reliability of the UK Biobank and ABCD phenotypes, the inverse-variance-weighted within-family association of the PGS with latent general ability is estimated at 0.448 (SE 0.025). Our PGS predicts higher educational attainment, occupational status, and family income within families, and retains good performance in non-European ancestry samples, with the correlation retaining 64% of its magnitude in the African American subset of the ABCD cohort, in line with expectations. In UKB siblings, higher scores correlate with better self-reported health and lower risk of multiple cardio-metabolic diseases. Our study shows that it is possible to attain powerful within-family prediction of GCA using a PGS, enabling future research to clarify how cognitive ability influences health-related outcomes.
The chief focus of the paper is looking at two complicating factors in polygenic score (PGS) validation: 1) within-family comparisons (siblings), and 2) the role of measurement error. Their Table 1 gives the main results:
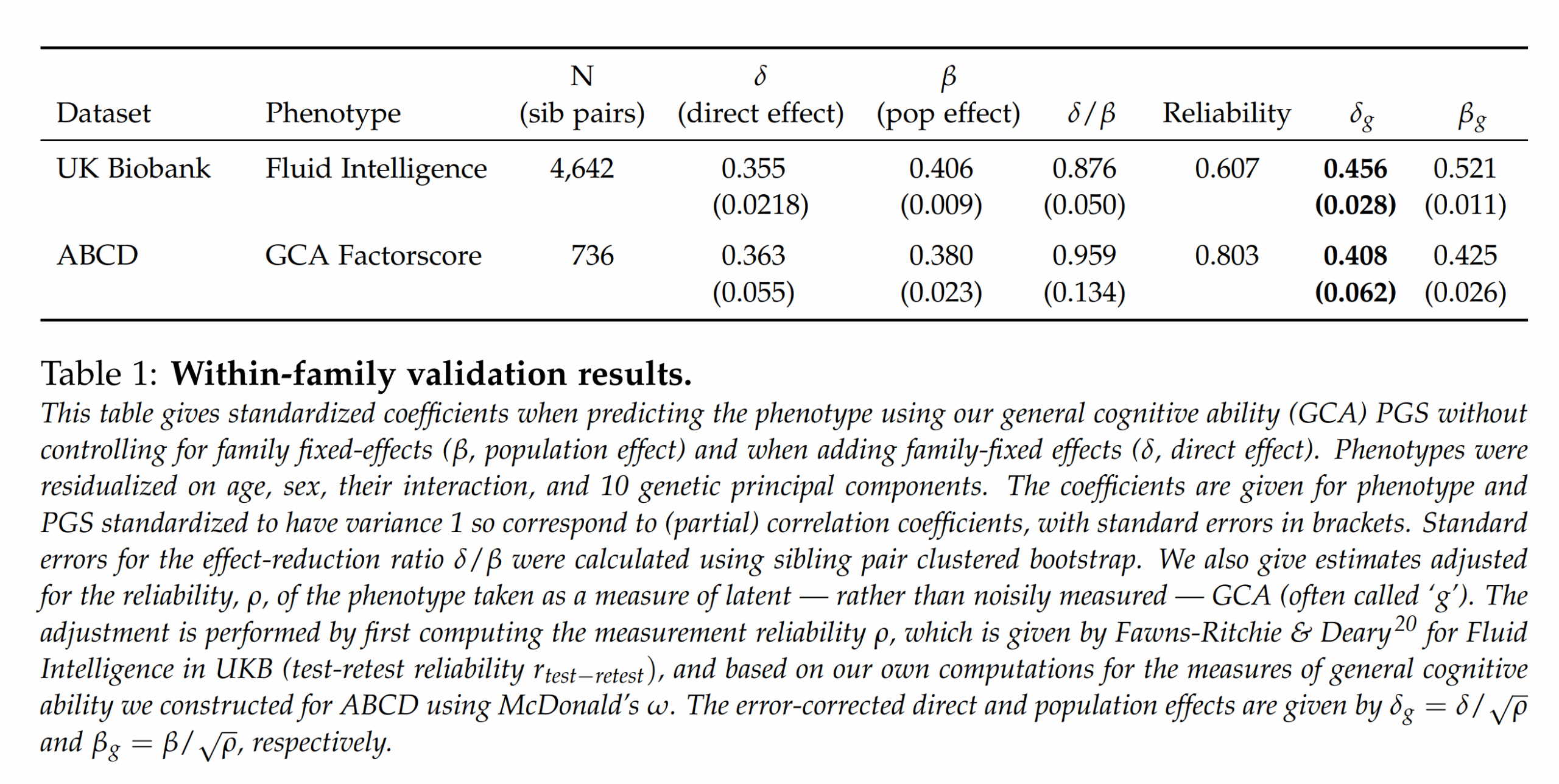
Using their proprietary model (based on SBayesRC), they achieve out of sample between family betas (correlation-metric) of about 0.40. Using sibling comparisons, they achieve about a beta of about 0.36. The ratios between the numbers are important here since it tells us how much of the validity works within family too. Since all embryo selection is choosing among siblings, between family validity is useless. Whenever a PGS works much less well within families than it does between, it suggests there is non-genetic confounding in the models, or something more complex like between-person gene-gene or gene-environment interactions or other interplay (e.g. genetic nurture; or parental genes that boost intelligence of children but only if they also share the same gene). The ratio they achieved is about 0.90, that is, 90% of the validity we see between families is also seen within families. Actually, given that the effect of measurement error is enhanced in sibling studies (because the model is relying on difference scores, each of which have their own error component which then is multiplied), this ratio is perhaps ~1 controlled for measurement error.
These are great results, but they go further and consider the classical test theory approach to PGS, something I’ve have been advocating for years. Given that the measures of intelligence in these samples is not perfect (it is never perfect), the unreliability (error) will cause the betas to be smaller than they really are. Thus, if we can estimate the size of this reduction via the reliability (the correlation of a test with itself taken again), then the true beta can be estimated. They did this to both of their samples. Here I want to nitpick about the UK Biobank data. They used just the fluid intelligence test (because it has the most data), which is quite poor, as it is only 13 items. You can in fact find the items here if you are curious. They are questions like “Add the following numbers together: 1 2 3 4 5 – is the answer?” and “If Truda’s mother’s brother is Tim’s sister’s father, what relation is Truda to Tim?” (with multiple choice). For this particular test, there is a subset of the data that took it twice, and the scores correlated at 0.607, the reliability seen in the table above. Dividing the betas by the square root of the reliability produces the estimated true betas, which were about 0.43 within family and 0.47 between (ratio 0.91; unweighted means). So what’s my complaint? The fluid reasoning test in the UKBB is rather limited as an intelligence test, and cannot be assumed to be perfectly loaded on g. Here’s a drawing to illustrate:
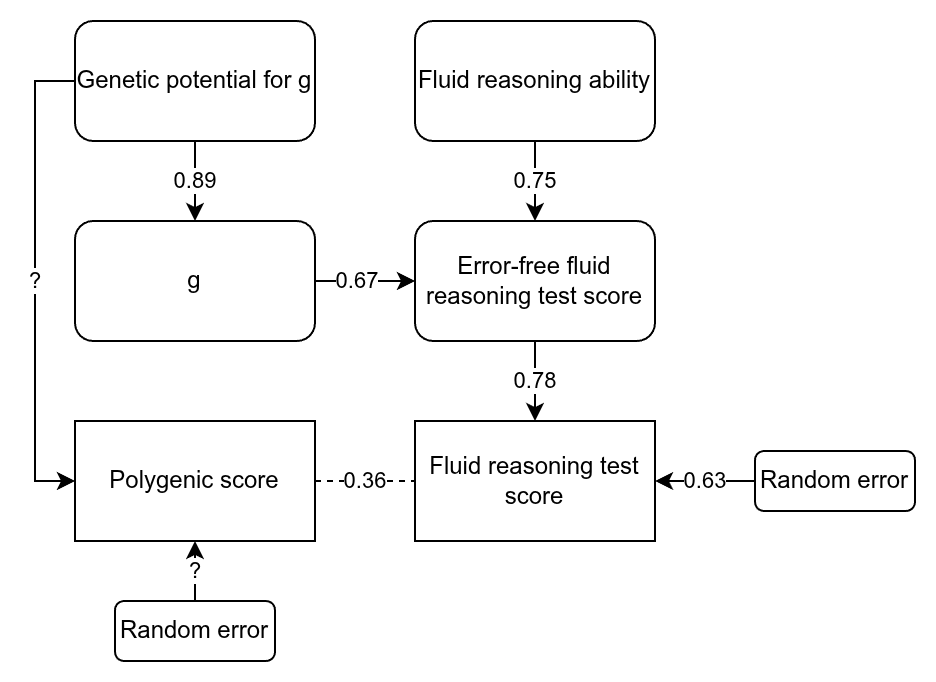
We can think of the raw score on the test as being caused by 3 variables we don’t observe directly: 1) general intelligence (g), 2) non-g ability related to fluid reasoning, and 3) random error. The study of the UKBB tests showed that the fluid test has a g-loading of 0.519. This is without taking into accounting measurement error. Using the quick method of adjusting this value for reliability (this assumes that the factor structure doesn’t change, which is not guaranteed given that tests have different amounts of error), gives a error-free g-loading of 0.67 = 0.519/sqrt(0.607). As such, the non-error other cause(s) of performance on this test must have a loading of 0.75. In other words, only about half the error-free variation in this test is due to g, the rest is other abilities, including prior experience (familiarity with similar content). I’ve also filled out some other paths, assuming a heritability of these samples of g of about 80% (their twin estimate for ABCD was 64%, which when corrected becomes 80%). So it means that when they find that they can predict the error-free variation in this test with a beta of 0.46 or so, this is a mixture of g and non-g factors. This problem does not occur in the ABCD because it has a battery of 10 tests of varying nature, meaning that the general factor of this battery is very close to being just g + error. This may explain some of the incongruence in their results, namely, that the error-free population effect sizes do not overlap in their confidence intervals. The error-free estimates for the direct effects overlap but have larger standard errors. Note that non-g abilities can also be quite heritable and more so when error is removed statistically, so just because they predict something heritable it doesn’t mean it is just g.
Next they consider the effect of error on heritability estimates using the all-snps approach, nowadays done using LDSC and before that using GREML. Since this is variance, the correction is to divide by the reliability itself, not the square root of it. They do simulations to show that this method works correctly. Strangely, they don’t appear to actually apply this to their h2_snp estimates. However, they do report that the UKBB h2_snp for the fluid test is 23%, which when corrected becomes 38%. They seemingly did not report the h2_snp for the ABCD. There are many studies analyzing the ABCD, maybe one of them has reported this value (this study has a lot of complex modeling but seemingly not the LDSC estimate). Anyway, this h2_snp value is an estimate of an additive, linear polygenic score trained on this dataset using these snps but with infinite sample size. The actual h2_pgs, that is, the heritability explained by the PGS is about 0.43^2=18%. As such, we could say that the current PGS captures about half the variance possible given the constraints (these particular genetic variants). However, practical utility for selection is a function of the beta, not the r²/h² values, so the PGS already captures about 70% of the validity (0.43/sqrt(0.38)).
Moving on to the strangest part about the study:
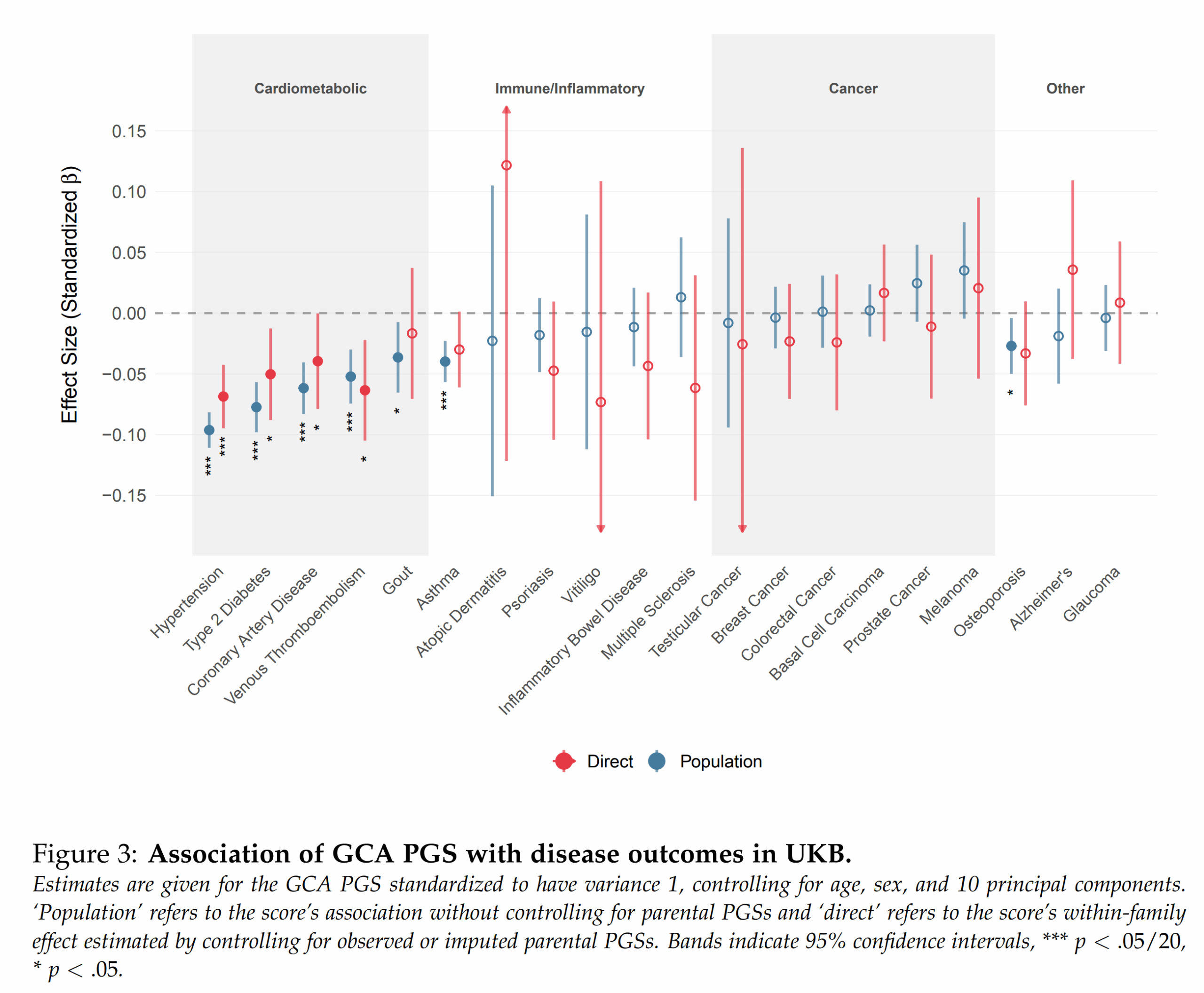
These are the particular validities of their g polygenic score for other outcomes, mainly diseases. In general, it is known that intelligence is mildly protective against a variety of diseases and that this effect is mainly genetic in origin. As such, selecting for higher intelligence gives one freebies on health traits. The strange thing here is that Alzheimer’s disease shows just about no effect. This is pretty strange considering that large national studies show that intelligence is protective in particular against this particular disease, and thus so should any PGS for intelligence.
They also looked at the other psychological measures in the ABCD and their intelligence PGS:
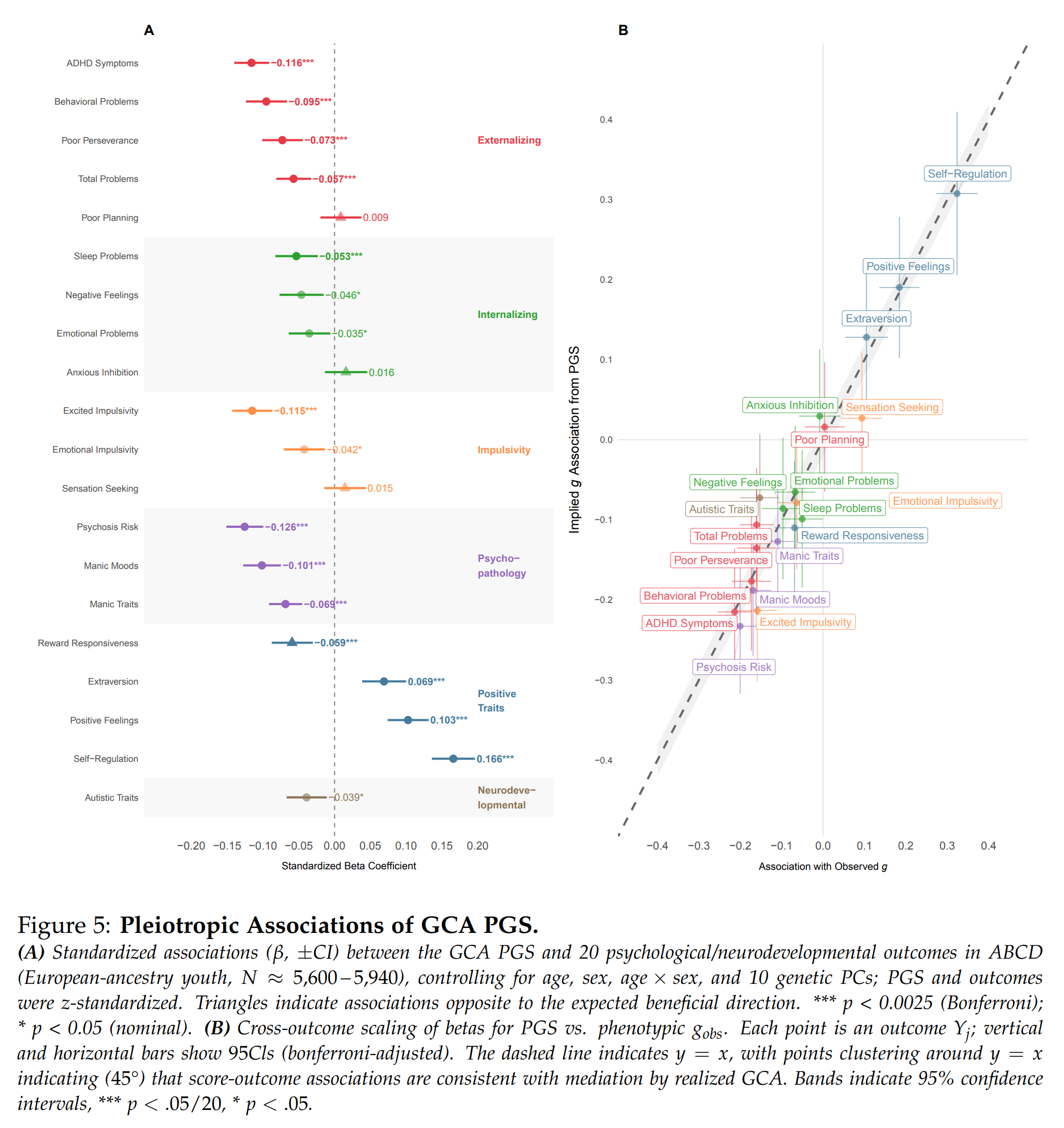
Since the genetic basis of many traits overlap to some extent (genetic correlations), selecting for one trait also means selecting, in part, for and against some of the others. Whether this is good or not depends on the directions of these correlations and how we value the traits. In this case, most people prefer not to have ADHD or their children to have it, and fortunately, the correlation is negative. The correlations for other mental problems were also mostly negative. There were positive correlations for ‘positive traits’, which includes extroversion. As such, selecting for intelligence using this score would also give you slightly more extroverted children. In this sample, their g PGS also had a slight negative correlation with autism symptoms. This is curious because many studies find small positive genetic correlations between PGSs for intelligence/education and autism. Maybe this is because this is a children’s sample, or because autism got more popular with the youth, thus making it less elite. Who knows.
Moving on. The other new embryo selection startup Nucleus Genomics has decided to publish their disease predictor models. This is an unexpected move towards open source genomics, and it’s most welcome. Actually, it’s one of those fake open access deals where you have to fill out an application. OK, let’s try! It’s at least less onerous than the academic equivalents. An improvement still, maybe.
They have trained these models on the UKBB and All of Us datasets I think, giving a total of maybe ~1.5M sample size. This means that Herasight and other parties can now (in theory) download these models and test them against their own models. Better yet, someone should set up an independent genetic model validation institution. It should collect a new sample measure their traits. Using these, it should run in-house validations of published genetic models. This way no one can cheat and overfit the models, and this institute can provide everybody with unbiased validation results, in the same way we have 3rd parties ranking existing AIs on a private IQ test the AIs have never seen. Many countries are building or expanding their biobanks, so it is not particularly difficult for them to set aside a few thousand people for such an institute’s internal dataset. If a few countries did this, the internal validation dataset could quickly reach 10k, including siblings for the crucial sibling-validation studies. What’s not to like about this idea? Maybe someone reading this blogpost can try to move forward with this idea. USA, for instance, is collecting a lot of data for their Million Veterans Project. This dataset is not used much by academics due to security clearance issues and paranoia of the military. However, team Trump could in theory just tell the military to do better, and even provide data for a genetic prediction model ‘clearinghouse’ in the public interest.