Sasha Gusev is a statistical geneticist who spends a lot of time on Twitter. He has been attacking various behavioral genetics studies lately, thus earning some ire from people in the field. Amusingly, he is seemingly unable to answer simple questions about his claims. I have so far asked him the same pertinent question 6 times. Each time he evades. The question concerns plots of polygenic score (PGS) averages vs. phenotypic averages (phenotype = the observed state of affairs). Most of my readers are probably familiar with Davide Piffer’s results, but here they are again from the 2021 paper:
Height polygenic vs. phenotypic scores in the 1000 genomes dataset:
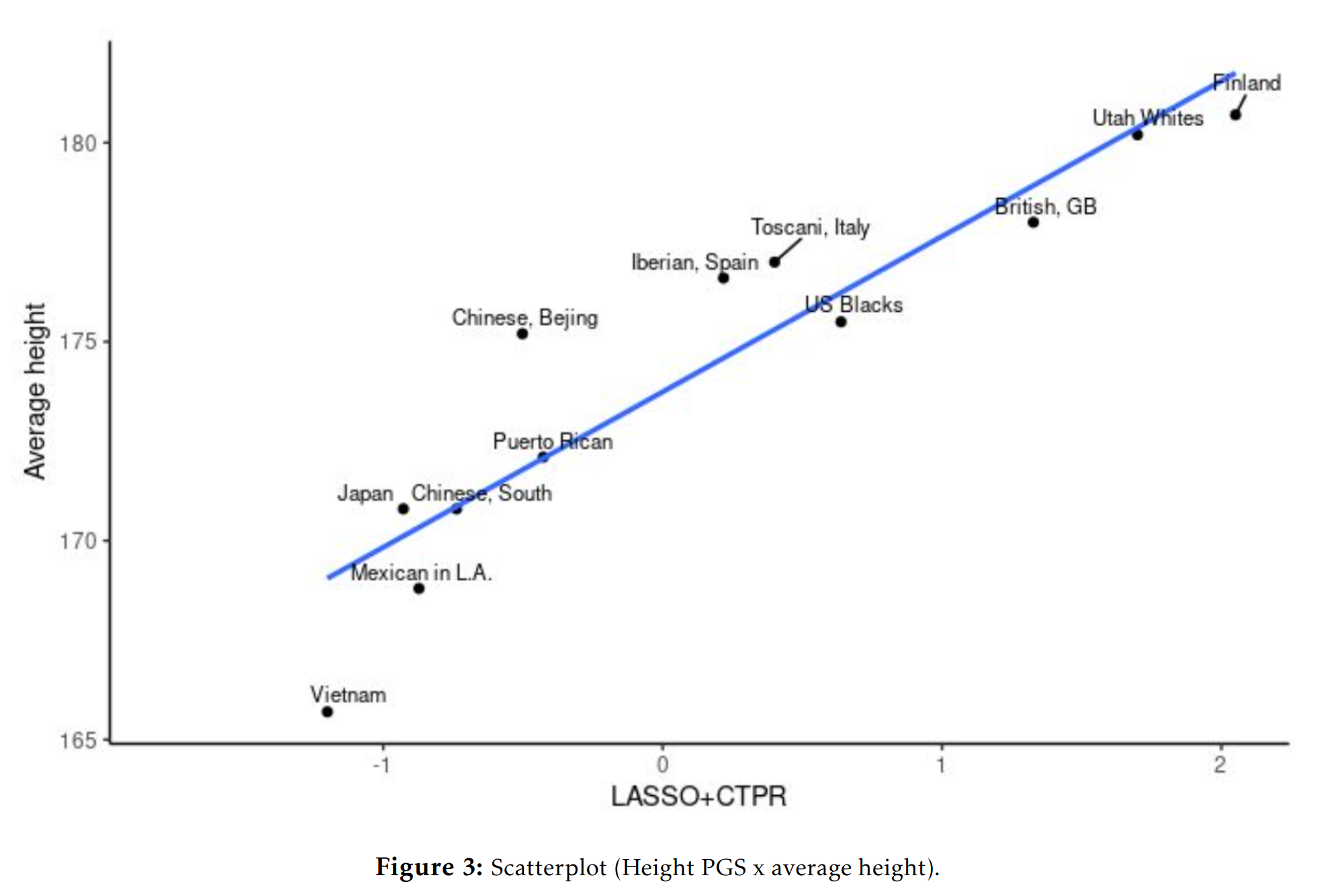
And in the GNOMAD dataset:
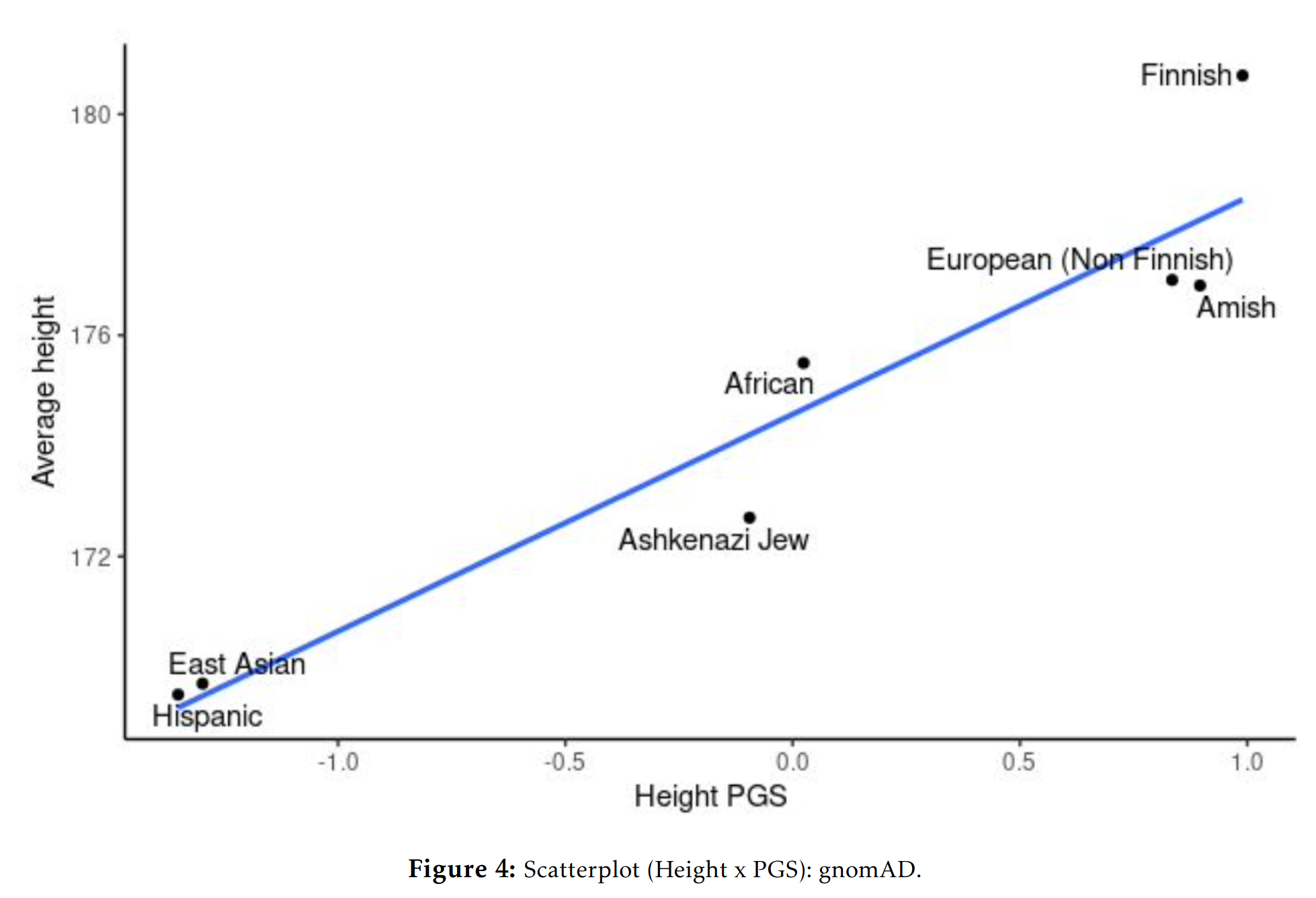
Education polygenic scores vs. phenotypic scores in the 1000 genomes dataset:
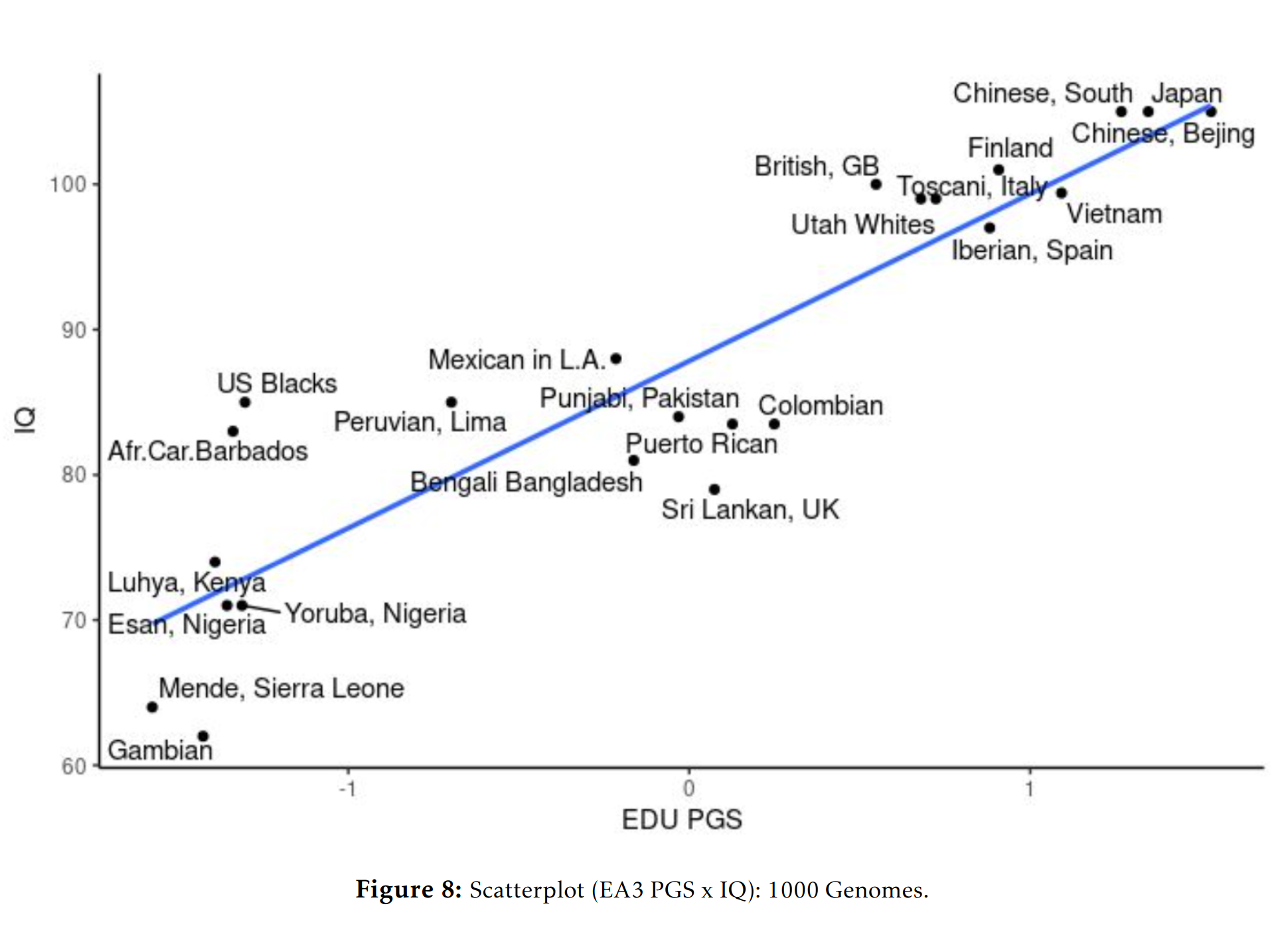
Education polygenic scores vs. phenotypic scores in the GNOMAD dataset:
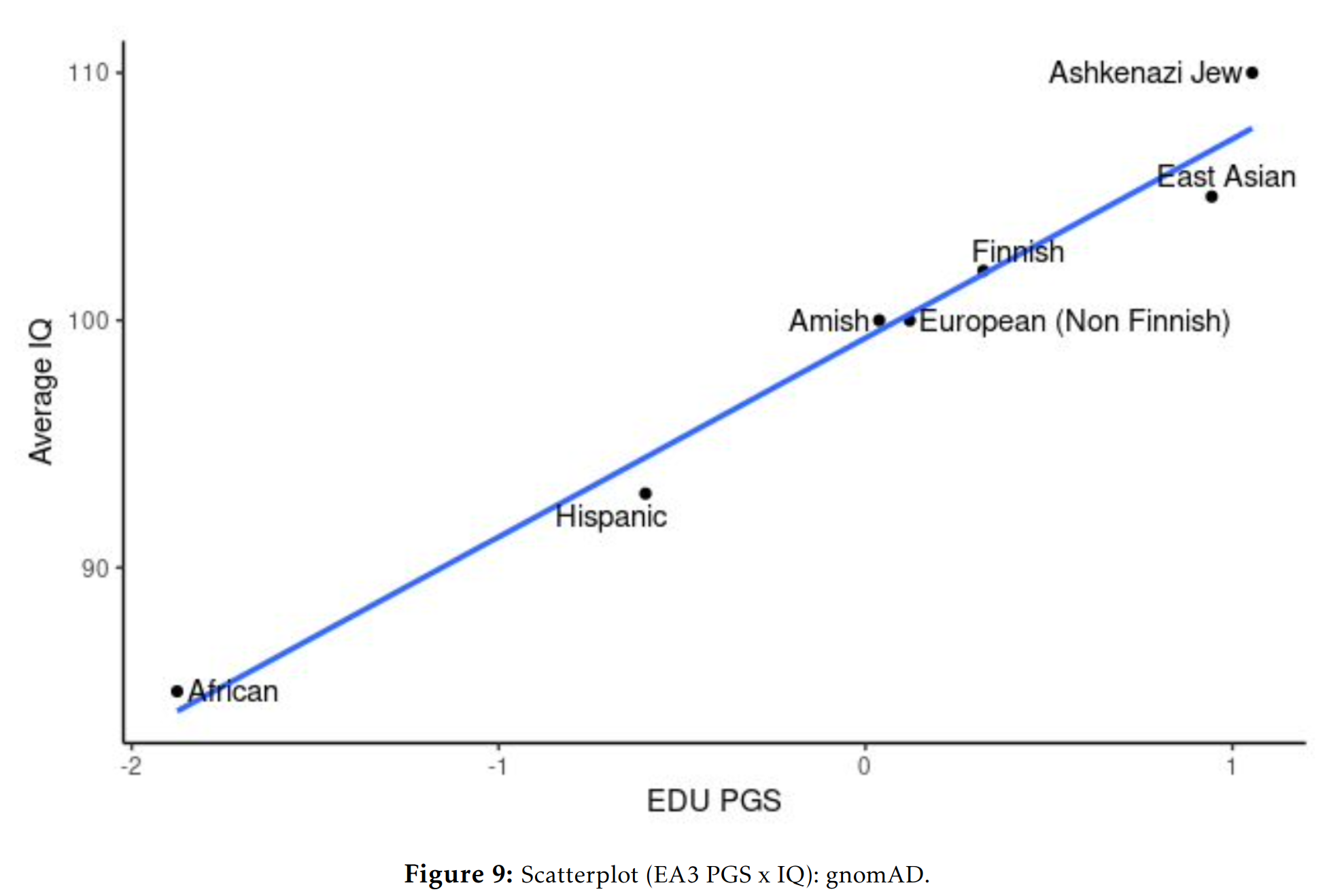
Education polygenic scores vs. phenotypes in the ABCD dataset, all of which are American children aged 10, from Fuerst et al 2023:
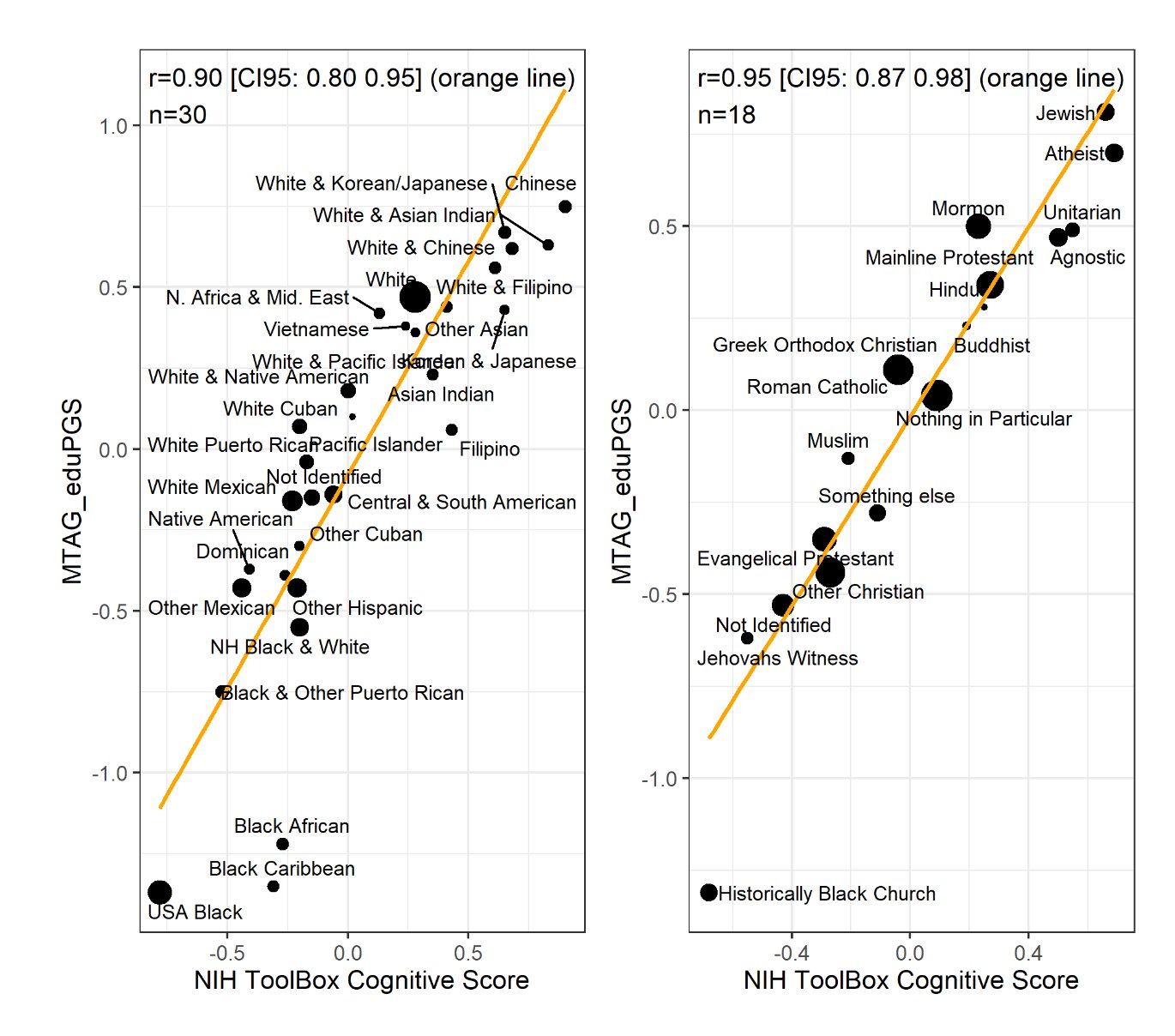
One could give more examples for other phenotypes, but the central question for any egalitarian is: why do the polygenic score means match up so well with the phenotypic means? Not perfectly, but decently given the current state of the art of polygenic scores. This question isn’t difficult from a hereditarian perspective: they line up reasonable well because the polygenic scores reflect the existing genetic differences between the populations’ genetic dispositions on various phenotypes and these are mostly but not entirely responsible for various group differences we see. There is no mystery. The models were trained only on Europeans with various controls for ancestries, and yet they keep lining up with the phenotypes. Why do they do so, if they don’t reflect a real signal? No one claims the current polygenic scores are perfect, or without bias. Hereditarians have been talking about these problems since the very first GWAS on education in 2013, I wrote about it in detail in 2015.
But Sasha Gusev just wont give us an answer:
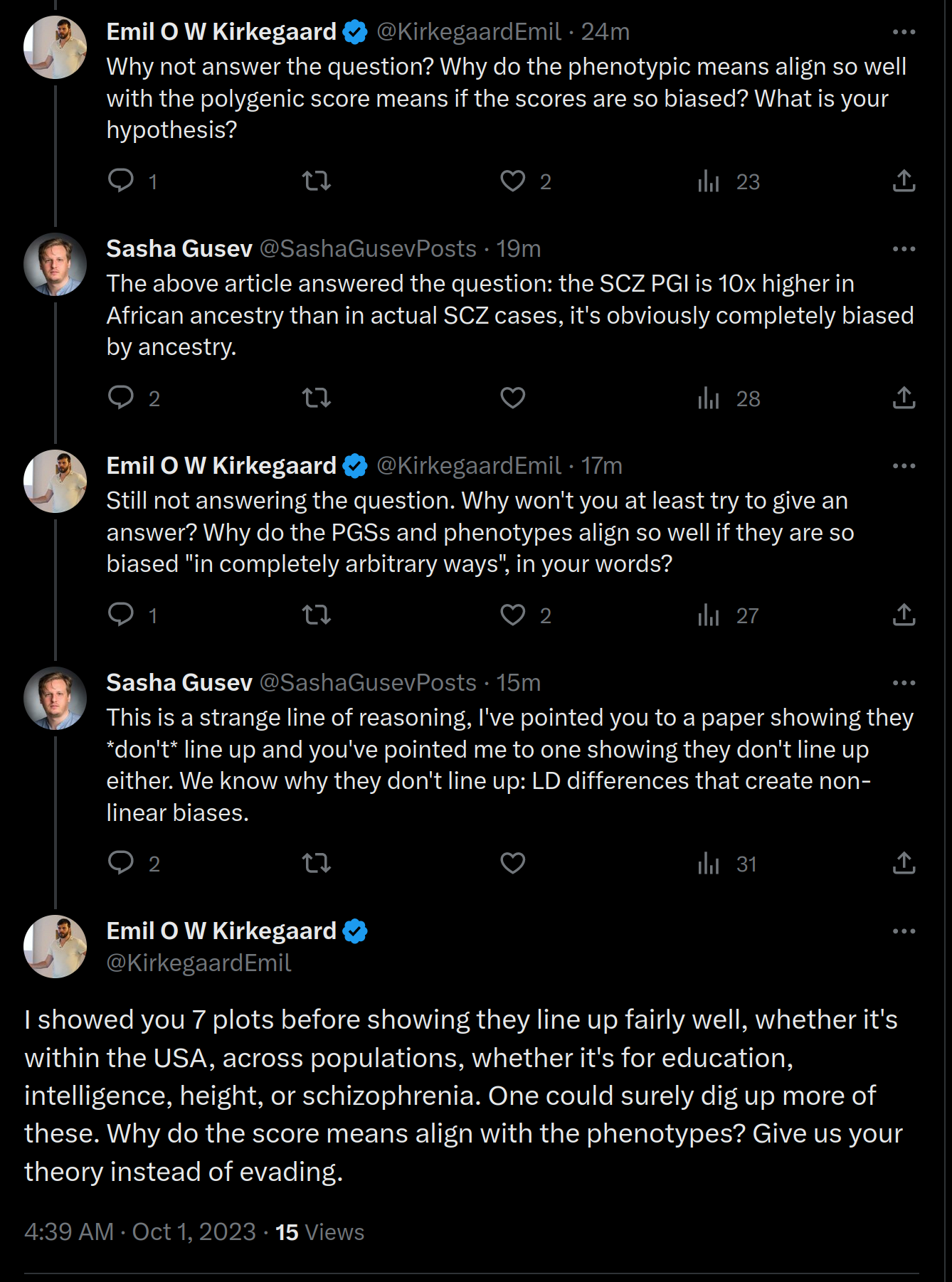
In fact, I haven’t seen any egalitarian explain their counter-model so far. They don’t seem to have one. Their attacks consist of doing weak selection tests (e.g. Kevin Bird), or pointing out that polygenic scores aren’t equally predictive across ancestries (yes, we know, but it’s not necessarily a problem). Based on this, I reckon they have two options. First, claim that the results are simply due to chance. As the results keep replicating with larger and more controlled GWASs, and larger samples of populations (we have 50+ in our working dataset), this answer becomes harder to maintain. Sasha Gusev seems to favor this hypothesis because as he claims:
I literally posted a screenshot of a European PGS mean changing in completely arbitrary ways across genetic distance with no relationship to the true trait mean. Not much of a reader?
Yes, I am aware one can make polygenic scores behave strangely. Yet, all the results above are from scoring them without any tricks, literally just out of the box. Why do the scores align so well?
Second, a better hypothesis is that the between family based GWASs still have some very subtle but important biases. A few years ago, there were a number of papers advocating this model, even going to far as to questioning whether height gaps were genetic in origin. They say to use sibling models, which by their nature have almost perfect controls “population stratification”, i.e., mistaking non-causal ancestry related genetic variation for causal genetic variation. A later sibling GWAS for height was able to reverse this conclusion again:
We found strong evidence for polygenic adaptation on taller height in the European meta-analysis GWAS using both population (r = 0.022; 95% CI 0.014, 0.031) and within-sibship GWAS estimates (r = 0.012; 0.003, 0.020) (Extended Data Figs. 5 and 6). These results were supported by several sensitivity analyses: (1) evidence of enrichment for positive tSDS (mean = 0.18, s.e. = 0.06, P = 0.003) amongst 310 putative height loci from the within-sibship meta-analysis results (Extended Data Fig. 7); (2) positive LDSC rg between height and tSDS in the meta-analysis results (Supplementary Table 8); and (3) evidence for polygenic adaptation on taller height when meta-analyzing correlation estimates from eight individual studies (for example, SDS using only UK Biobank GWAS summary data) for population (r = 0.013; 0.010, 0.015) and within-sibship (r = 0.004; 0.002, 0.007) estimates (Fig. 7). There was also some putative within-sibship evidence for polygenic adaptation on increased number of children (P = 0.024) and lower high-density lipoprotein (HDL)-cholesterol (P = 0.024) (Extended Data Fig. 5).
Hereditarians expect, of course, that as sibling GWASs become larger and larger, we will start seeing evidence for selection for other phenotypes. Height is the easiest phenotype to show selection for because the gaps are large, not controversial, and easy to measure well. On the other hand, differences in intelligence are harder to measure and politically fraught. The same thing goes for differences in, say, schizophrenia.
For the purpose of this blogpost, I redid the plots from Davide Piffer’s recent 2023 article (data here) using the new sibling GWAS results for education (Howe 2022):
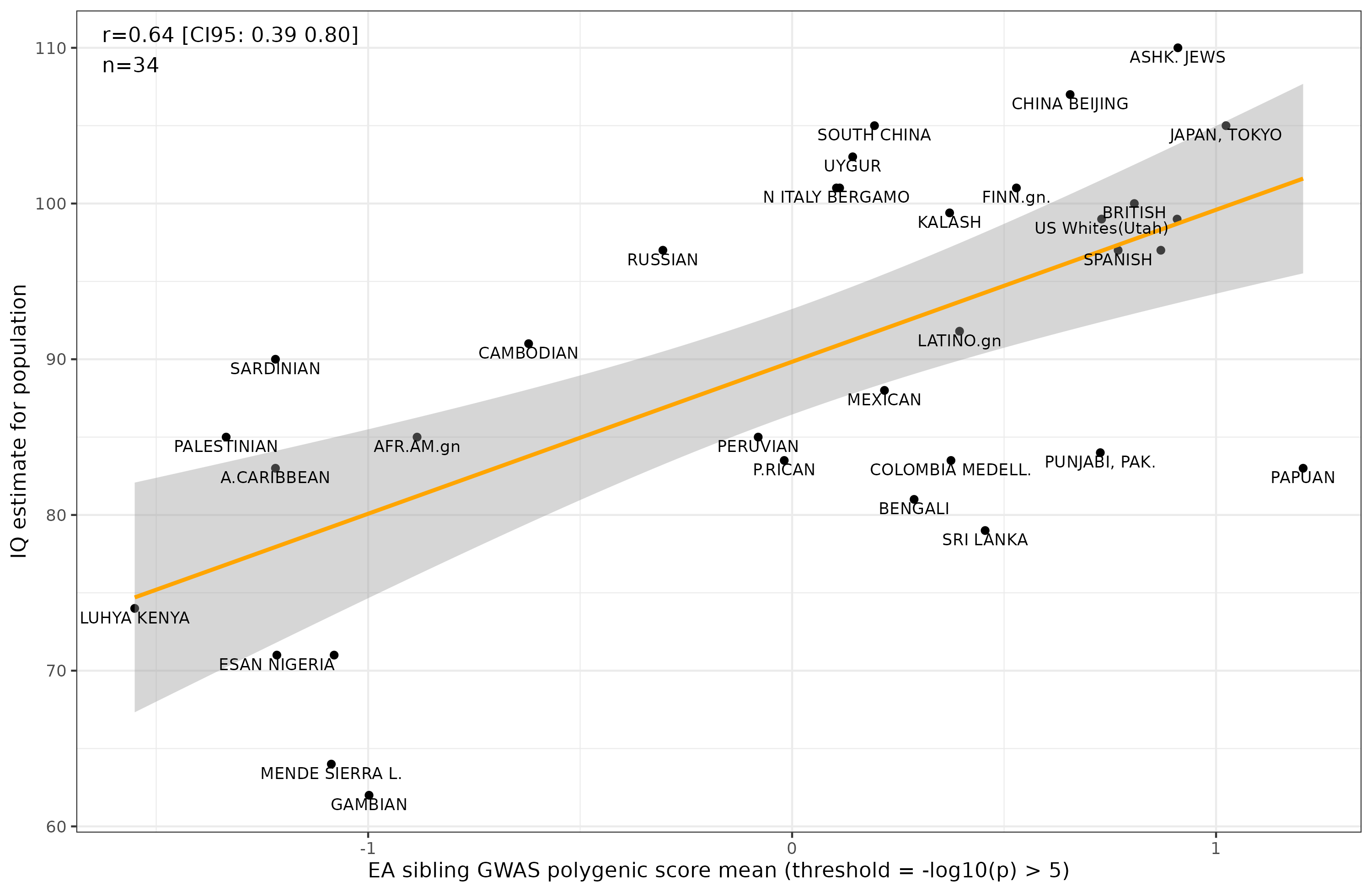
It’s a bit less impressive than the much larger EA4 education between-family GWAS (Okbay 2022):
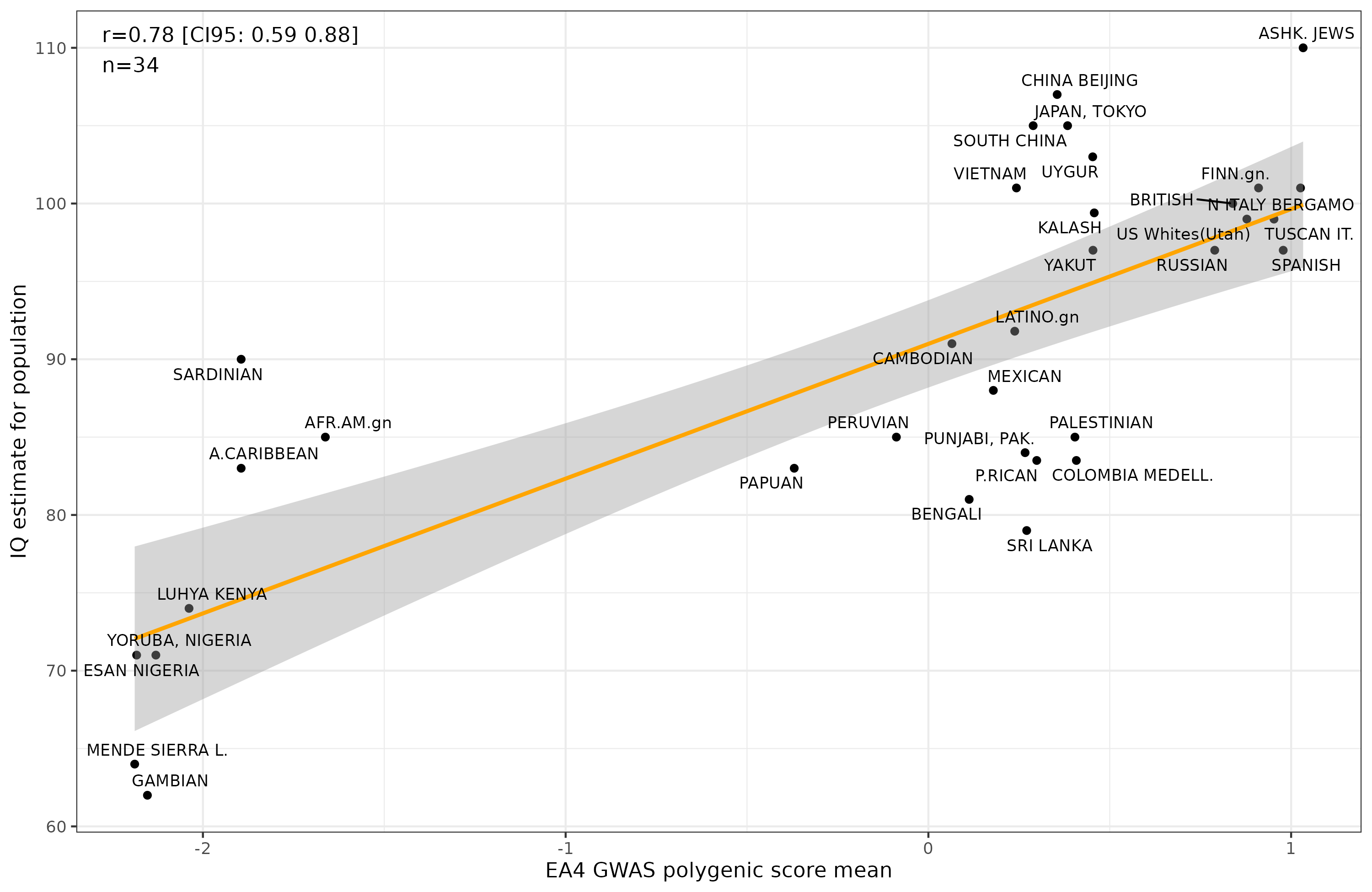
Still, the values aren’t too far off from each other, and both are p < .01. At this sample size (n=34 groups), one cannot really say whether these correlations are different from each other. We are spending a lot of time compiling more datasets so that more groups and countries can be included. Hopefully, we can get to n > 100 within a year.
Conclusions
The evidence from polygenic scores is suggestive, not final. The coming decade will reveal more clear evidence as trans-ethnic GWASs on education/intelligence will be done, larger sibling or other family GWASs, and GWASs based on whole genome sequencing (which removes the ascertainment bias in European-based array data). The case for hereditarianism does not rest on the current polygenic score results, as many egalitarians appear to believe. There are many summaries of the evidence — including my own from a few months ago — but in point form:
- Polygenic score gaps align with known phenotypes as expected.
- Admixture regression results suggest substantial genetic causation of European vs. African or vs. Amerindian gaps for intelligence, as based on detailed analyses of PING, PNC, ABCD datasets. Other researchers have also replicated these for social status many times in the last 10 years.
- Measurement invariance testing rules out various X factor theories, which together with approximately even heritabilities makes Jensen’s variance argument hard to get around. Simply speaking, the causes of group differences in intelligence are the same as those that cause within group differences, and the group gaps in social status just aren’t large enough to explain the intelligence gaps, even if one makes implausible assumptions about their size.
- The failure of 100s of billions of dollars spent on remedial, compensatory, early education the last 70 years suggest the causes aren’t environmental. In fact, the gaps have remaining basically stable for 100 years despite a complete change form legal anti-Black racism in the US south, to mandatory pro-Black racism in every state, and a related fall in anti-Black sentiment among Whites. If these factors are important, then why did the gap not change?
- There are positive correlations between items’ and tests’ g-loadings, heritabilities (family and polygenic scores), group differences based on social race, and ancestry associations. This total pattern is difficult to explain from an egalitarian perspective.
- Detailed analyses of proposed causes of intelligence gaps show that these fall woefully short. For instance, subclinical lead poisoning is often claimed, but rigorous research suggest tiny effect sizes of lead, and in any case, the group gaps on this variable aren’t large enough.